Output format
Graphical output
The output examples displayed on this page, was obtained by going to the SigniSite-2.1 server [1] and clicking 'Load sample data' followed by 'Submit' under the SigniSite logo.
This will generate the following figures
Figure 1: Sequence logo quantifying strength of residue association
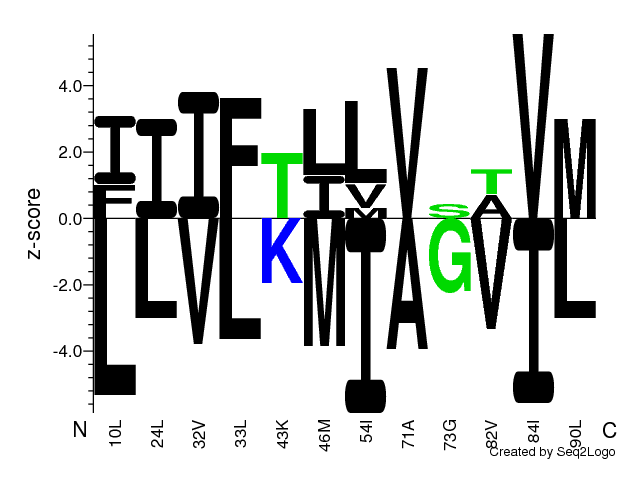
Logo quantifying strength of residue association [2].
Amino acid residues on the positive y-axis are associated with strong phenotype values
and residues on the negative y-axis, with weak phenotype values, i.e. residues above the
z=0.0 line have a z-score larger than zero and are thus predominantly found among the top
of the sorted aligned sequences. E.g. low binding affinities or high luminescence signals
delending on the users choice of sequence sorting. Vice versa for residues below the
z=0.0 line.
The amino acids are colored according to their chemical properties as follows:
Acidic [DE]: red, Basic [HKR]: blue, Hydrophobic [ACFILMPVW]: black and Neutral
[GNQSTY]: green. [3]. If any of the sites are denoted by negative numbers, this
implies that a reference sequence was chosen and these sites lie in gapped regions.
Gapped regions are regions, in which insertion (ins) is found. Please note that if
a reference sequence was chosen, this sequence will be given below each column in
the logo plot, rather than the consensus sequence.
-1 is the first ins, -2 the second ins and so on.
Figure 2: Heatmap visualisation of strength of residue association
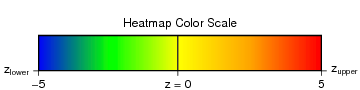
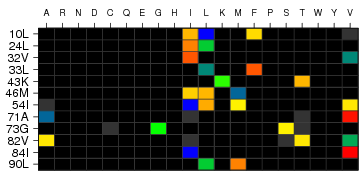
Heatmap visualisation of strength of residue association. The color-scale
(See 'Heatmap Color Scale') ranges from blue z < -5 to red z >5.
For z-scores larger than -5, but smaller than 5 colors inbetween are
used. Black cells denote abscence of amino acid residue. A grey cell denotes a residue
with a z-score of 0. If there is only one grey cell at a position, the position
is completely conserved harbouring only this residue. If more than one grey cell are
present, the p-value for this residue has become p = 1 after correction for
multiple comparisons. Each column corresponds to one of the 20 proteinogenic amino acids
and each row to a position in the submitted multiple sequence alignment. If any of the sites
are denoted by negative numbers, this implies that a reference sequence was chosen and these
sites lie in gapped regions. Please note that if a reference sequence was chosen, this
sequence will be given below each column in the logo plot, rather than the consensus sequence.
Gapped regions are regions, in which an insertion (ins) is found. -1 is the first ins, -2 the
second ins and so on.
Additional output
Other than the graphical output, the following output files will be available:
- Alignment
- - The multiple sequence alignment used for the analysis
- Excel file 1 (.csv with blanks)
- - Excel compatible z-score table. All non-present residues are blank
- Excel file 2 (.csv with zeros)
- - Excel compatible z-score table. All non-present residues are '0.000'
- HTML score table
- - Printer friendly z-score table in HTML format.
- Weight matrix/PSSM
- - Position Specific Scoring Matrix (PSSM)
- Rank list of z-scores
- - Ranked list of z-scores. Columns are:
- #Pos: The position in the submitted multiple sequence alignment
- Cons: The consensus residue at the position
- Resi: The amino acid residue for which the z-score was computed
- Asso: Positive or negative association of z-scores (denotes z<0 or z>0)
- Zsco: The absolute value of the computed z-score (i.e. 'ignoring' the sign of z)
- Pval: The p-value corresponding to the computed z-score
- Rank: The rank of the z-score (Tied values are assigned mean rank)
References
For publication of results, please cite [1]
-
Jessen LE, Hoof I, Lund O, Nielsen M.
SigniSite: Identification of residue-level genotype-phenotype correlations in protein multiple sequence alignments.
Nucleic Acids Res. 2013 Jul;41(Web Server issue):W286-91. doi: 10.1093/nar/gkt497. Epub 2013 Jun 12.
-
Thomsen MC, Nielsen M.
Seq2Logo: a method for construction and visualization of amino acid binding motifs and sequence profiles including sequence weighting, pseudo counts and two-sided representation of amino acid enrichment and depletion.
Nucleic Acids Res. 2012 Jul;40(Web Server issue):W281-7. doi: 10.1093/nar/gks469. Epub 2012 May 25.
-
Lund, O., Nielsen, M., Lundegaard, C., Kesmir, C., Brunak, S.
Immunological Bioinformatics.
(S. Istrail, P. Pevzner, M. Waterman, Eds.), (1st ed., p. 312). Cambridge, Massachusetts, London, England: The MIT Press. ISBN-10: 0262122804, ISBN-13: 9780262122801. Jul 2005


