DTU Health Tech
Department of Health Technology
This link is for the general contact of the DTU Health Tech institute.
If you need help with the bioinformatics programs, see the "Getting Help" section below the program.
DTU Health Tech
Department of Health Technology
This link is for the general contact of the DTU Health Tech institute.
If you need help with the bioinformatics programs, see the "Getting Help" section below the program.
The NetMHCIIpan-4.2 server predicts peptide binding to HLA class II molecules using Artificial Neural Networks (ANNs). It is trained on an extensive dataset of over 600.000 measurements of Binding Affinity (BA) and Eluted Ligand mass spectrometry (EL), covering the three human MHC class II isotypes HLA-DR, HLA-DQ, HLA-DP, as well as mouse molecules (H-2).
The network can predict for any HLA class II molecule of known sequence, which the user can specify as FASTA format, and predictions can be made for peptides of any length.
The output of the model is a prediction score for the likelihood of a peptide to be naturally presented by an MHC-II receptor of choice. The output also include a %rank score, which normalizes the prediction score by comparing to predictions of a set of random peptides. Optionally, the model also outputs BA prediction and %rank scores.
New in version 4.2: The method is trained on a extended set of HLA-DQ EL data compared to NetMHCIIpan-4.1.
Refer to the instructions page for more details.
The project is a collaboration between DTU-Bioinformatics, and LIAI.
View the version history of this server
UPDATE (September 27, 2023): Some missing pre-calculated files to estimate percentile ranks, along with the option to include BA predictions, have been added to the webserver and downloadable executables. If you downloaded NetMHCIIpan-4.2 before this date, please redownload the package (4.2c) to get the updated version.
NetMHCIIpan 4.2 is available as a stand-alone software package, with the same functionality as the service above. Ready-to-ship packages exist for Linux and macOS. There is a download page for academic users; other users are requested to contact Health Tech Software Package Manager at health-software@dtu.dk.
In this section, the user must define the input for the prediction server following these steps:
1) Specify the desired type of input data (FASTA or PEPTIDE ) using the drop down menu.
2) Provide the input data by means of pasting the data into the blank field, uploading it using the "Choose File" button or by loading sample data using the "Load Data" button. All the input sequences must be in one-letter amino acid code. The alphabet is as follows (case sensitive):
A C D E F G H I K L M N P Q R S T V W Y and X (unknown)
Any other symbol will be converted to X before processing. At most 5000 sequences are allowed per submission; each sequence must be not more than 20,000 amino acids long and not less than 9 amino acids long.
3) If FASTA was selected as input type, the user must select the peptide length(s) the prediction server is going to work with. NetMHCIIpan-4.2 will "chop" the input FASTA sequence in overlapping peptides of the provided length and will predict binding against all of them. By default input proteins are digested into 15-mer peptides. Note that, if PEPTIDE was selected as input type, this step is unnecessary and thus the peptide length selector will directly not appear in the interface.
4) Context encoding informs the network of the proteolytic context the ligand. Context is automatically generated from the source protein if the user selects FASTA format. Briefly, context is made up of 12 amino acids: 3 amino acids upstream of the ligand, 3 first amino acids at the ligand N-terminus, 3 last amino acids at the ligand C-terminus and 3 amino acids downstream the ligand(in the source protein), all concatenated together. If the input type is PEPTIDE , the user must specify the ligand context(see PEPTIDECONT ).

In this section, the user must define which MHC molecule(s) the input data is going to be predicted against:
1) Here the user can select from a list of MHC molecules by first selecting the group and clicking MHCs in the list. Here, MS-COVERED refers to molecules properly covered by the NetMHCIIpan-4.2 training data.
2) The user can also type the molecule names. Both the ALPHA and BETA chains must be typed (please consult List of MHC molecule names.) Note that molecules selected from step 1 populate this bar.
3) If the molecule of interest is not provided in the lists, the user can input ALPHA and BETA sequences in fasta format. With this option, rank score predictions are not available.

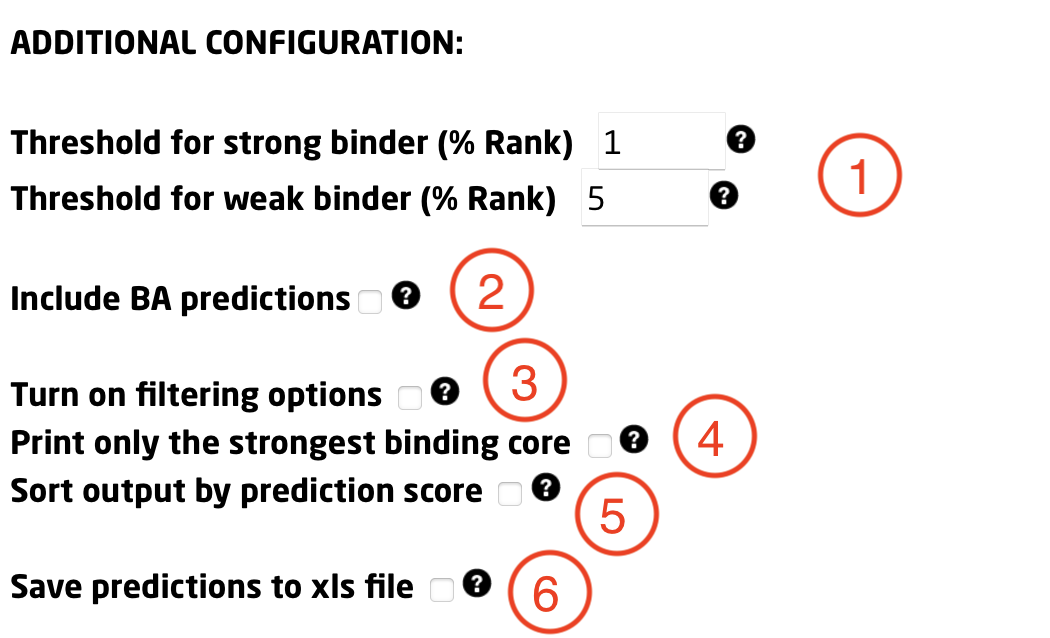
In this section, the user may define additional parameters to further customize the run:
1) Specify thresholds for strong and weak binders. They are expressed in terms of %Rank, that is percentile of the predicted binding affinity compared to the distribution of affinities calculated on set of random natural peptides. The peptide will be identified as a strong binder if it is found among the top x% predicted peptides, where x% is the specified threshold for strong binders (by default 2%). The peptide will be identified as a weak binder if the % Rank is above the threshold of the strong binders but below the specified threshold for the weak binders (by default 10%).
2) Tick this option to include also Binding Affinity predictions together with Eluted Ligand likelihood.
3) Tick this option to output only peptides with a % Rank score below a specified threshold. Useful for large submissions.
4) Tick this box to output only the strongest binding core.
5) Tick this box to have the output sorted by descending prediction score.
6) Enable this option to export the prediction output to .XLS format (readable for most spreadsheet softwares, like Microsoft Excel).

# NetMHCIIpan version 4.2 # Input is in FASTA format # Peptide length 15 # Prediction Mode: EL # Threshold for Strong binding peptides (%Rank) 1% # Threshold for Weak binding peptides (%Rank) 5% # DQA10301-DQB10302 : Distance to training data 0.0000 (using nearest neighbor HLA-DQA10301-DQB10302) # Allele: DQA10301-DQB10302 -------------------------------------------------------------------------------------------------------------------------------------------- Pos MHC Peptide Of Core Core_Rel Identity Score_EL %Rank_EL Exp_Bind BindLevel -------------------------------------------------------------------------------------------------------------------------------------------- 3 DQA10301-DQB10302 EMKTDAATLAQEAGN 4 DAATLAQEA 0.590 P9WNK5 0.467401 0.23 NA <=SB 78 DQA10301-DQB10302 QAGVQYSRADEEQQQ 3 VQYSRADEE 0.900 P9WNK5 0.418179 0.35 NA <=SB 4 DQA10301-DQB10302 MKTDAATLAQEAGNF 3 DAATLAQEA 0.770 P9WNK5 0.406498 0.38 NA <=SB 2 DQA10301-DQB10302 AEMKTDAATLAQEAG 5 DAATLAQEA 0.530 P9WNK5 0.381155 0.47 NA <=SB 77 DQA10301-DQB10302 RQAGVQYSRADEEQQ 4 VQYSRADEE 0.890 P9WNK5 0.348975 0.64 NA <=SB 5 DQA10301-DQB10302 KTDAATLAQEAGNFE 2 DAATLAQEA 0.790 P9WNK5 0.325364 0.79 NA <=SB 79 DQA10301-DQB10302 AGVQYSRADEEQQQA 2 VQYSRADEE 0.740 P9WNK5 0.295187 1.03 NA <=WB 1 DQA10301-DQB10302 MAEMKTDAATLAQEA 6 DAATLAQEA 0.420 P9WNK5 0.260940 1.41 NA <=WB 76 DQA10301-DQB10302 IRQAGVQYSRADEEQ 5 VQYSRADEE 0.960 P9WNK5 0.250306 1.54 NA <=WB 80 DQA10301-DQB10302 GVQYSRADEEQQQAL 1 VQYSRADEE 0.600 P9WNK5 0.160670 3.56 NA <=WB
The prediction output for each molecule consists of the following columns:
Jonas Birkelund Nilsson, Saghar Kaabinejadian, Hooman Yari, Bjoern Peters, Carolina Barra, Loren Gragert, William Hildebrand and Morten Nielsen
Here, you will find the data set used for training of NetMHCIIpan-4.2.
Download the file and untar the content using
cat NetMHCIIpan_train.tar.gz | tar xvf -
This will create the directory called NetMHCIIpan_train. In this directory you will find 12 files. 10 files (c00?_ba, c00?_el) with partitions with binding affinity (ba) with eluted ligand data (el). The format for each file is (here shown for an el file)
AAAAAAAAAAAA 1 Saghar_9075 AGRAAAAAAAAA AAAAAAAAAAAA 1 Saghar_9090 AGRAAAAAAAAA AAAAAAAAAAAAA 1 Bergseng__9037_SWEIG AGRAAAAAAAAG AAAAAAAAAAAAA 1 Bergseng__9064_AMALA AGRAAAAAAAAG AAAAAAAAAAAAA 1 Bergseng__9089_BOB AGRAAAAAAAAG AAAAAAAAAAAAA 1 Saghar_9052 AGRAAAAAAAAG AAAAAAAAAAAAA 1 Saghar_9075 AGRAAAAAAAAG AAAAAAAAAAAAA 1 Saghar_9090 AGRAAAAAAAAG AAAAAAAAAAAAAA 1 Bergseng__9037_SWEIG AGRAAAAAAAGA AAAAAAAAAAAAAA 1 Bergseng__9064_AMALA AGRAAAAAAAGAwhere the different columns are peptide, target value, MHC_molecule/cell-line, and context. In cases where the 3rd columns is a cell-line ID, the MHC molecules expressed in the cell-line are listed in the allelelist.txt file.
The allelelist.txt file contains the information about alleles expressed in each MA cell line data set, and pseudosequence.2016.fix the MHC pseudo sequenes for each MHC molecule.
Please click on the version number to activate the corresponding server.
| 4.2 |
The current server (online since September 2022). New in this version:
|
| 4.1 |
(online since Sept 2021). New in this version:
|
| 4.0 |
(online since April 2020). New in this version:
|
| 3.2 |
(online since January 2018). New in this version:
|
| 3.1 |
(online since December 2014). New in this version:
|
| 3.0 |
(online since June 2013). New in this version:
|
| 2.1 |
(online since 6 June 2011). New in this version:
|
| 2.0 |
(online since 17 Nov 2010). New in this version:
|
| 1.1 |
(online since 15 April 2010). New in this version:
|
| 1.0 |
Original version (online version until April 15 2010):
Main publication:
|
If you need help regarding technical issues (e.g. errors or missing results) contact Technical Support. Please include the name of the service and version (e.g. NetPhos-4.0) and the options you have selected. If the error occurs after the job has started running, please include the JOB ID (the long code that you see while the job is running).
If you have scientific questions (e.g. how the method works or how to interpret results), contact Correspondence.
Correspondence:
Technical Support: