 Click image for larger version
Click image for larger version
| Target MIM: | 151623, LI-FRAUMENI SYNDROME 1; LFS1 |
| Number of genes in interval: | 335 |
| Posterior probability for the gene: | 1.000 |
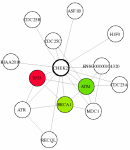
| Gene: | CHEK2 (ENSG00000183765) |
Read the cytoscape guide for instructions on how to use these files.
Download the cytoscape graphical properties file for this paper: lage_karlberg_et_al.props
| ENSG00000183765.sif | Network topology specification. |
| ENSG00000183765_input.noa | Node attribute file. Sets the attribute "input" to 1 for the candidate protein. |
| ENSG00000183765_labels.noa | Node attribute file. Specifies the gene name for each node. |
| ENSG00000183765_description.noa | Node attribute file. Functional description from Ensembl for each gene(node) |
| ENSG00000183765_mims.noa | Node attribute file. Lists any MIMs associated with each protein and the phenotype association score between the these MIMs and the target MIM |
| ENSG00000183765_associations.noa | Node attribute file. Contains the best phenotype association score between the target MIM and any phenotype to which the gene(node) is associated. |
| ENSG00000183765_score.eda | Edge attribute file. Interaction score for the protein protein interaction |
| ENSG00000183765_support.eda | Edge attribute file. Supporting sources for the protein protein interactions in the form of a list separated by '::'. Each element in the list is further separated with '|' and the fields are 'accession A|accession B|source db|interaction representation type||pubmed id' |
| 151623-ENSG00000183765.zip | All of the above files in a zip archive |