DTU Health Tech
Department of Health Technology
This link is for the general contact of the DTU Health Tech institute.
If you need help with the bioinformatics programs, see the "Getting Help" section below the program.
DTU Health Tech
Department of Health Technology
This link is for the general contact of the DTU Health Tech institute.
If you need help with the bioinformatics programs, see the "Getting Help" section below the program.
The NNAlign server allows generating artificial neural network models of receptor-ligand
interactions. The program takes as input a set of ligand sequences with target values; it returns
a sequence alignment, a binding motif of the interaction, and a model that can be used to
scan for occurrences of the motif in other sequences.
Visit the links on the pink bar below to read detailed instructions and guidelines, see output formats,
or download the code.
New in version 2.0/2.1:
For publication of results, please cite:
Job name:
This prefix is pre-pended to all files generated by the current run. If left empty, a
system-generated number will be assigned as prefix.
Motif length:
The length of the alignment core can be specified as:
Order of the data:
By default, peptides with high values are positive instances and high prediction scores are used to derive the sequence logo. You can invert this behaviour
Data rescaling:
The optimal data distribution for NN training is between 0 and 1 with the bulk of the data in the middle region of the spectrum. With the default option the program rescales linearly the data between 0 and 1, but it is also possible to apply a logarithmic transformation if the data appears squashed towards lower values. If your data is already rescaled between 0 and 1, select the No rescale option
You can also inspect the data distribution before and after the transformation in the output panel, following the link "View data distribution". Example
Average target values of identical sequences:
If there are duplicated sequences, by default they are all used together with their target values. Toggle this option to only use each sequence once with the average of the multiple target values.
Folds for cross-validation:
Specify the number of subsets to be created for the estimation of performance on
cross-validation. It is also possible to skip cross-validation, ticking the 'NO' button. In this case all data are used for training, and execution will be faster, but it won't be possible to calculate performance measures.
Cross-validation method:
The predictive performance of the method is estimated in cross-validation (CV) on the
training set. At each cross-validation step, one of 'n' subsets is left out as an evaluation set, where 'n' is the number of folds, rotating the evaluation set n times. Two CV methods are available:
Stop training on best test-set performance:
If this option is selected, training of the networks will be stopped on the highest CV test-set performance (Early stopping). A completely unbiased evaluation of the performance requires an additional independent test set, by selecting Nested cross-validation. However, for large datasets an accurate and much faster estimate of the predictive performance can be done on the same subsets used for early stopping (Simple cross-validation together with Early stopping).
Leaving the Early stopping option unticked will continue the training until the maximum number of training cycles as specified in the "Number of training cycles" option.
Method to create subsets:
The data can be prepared for cross-validation in 4 manners:
Remove homologous sequences from training set:
Homologous sequences are by default clustered in the same subset. Check the box to keep only one instance of homologous sequences.
Alphabet:
You may use a custom alphabet (e.g. nucleic acids, or non-standard amino acids). All upper-case letters and the symbols + and @ are permitted. The symbol X is reserved as a wildcard. Note that if you modify the alphabet, all BLOSUM options will be disabled.
Number of training cycles:
This option specifies how many times each example in the training set is
presented to the neural networks. If training is stopped on the best test-set performance,
this value represents the maximum number of training cycles.
Number of seeds:
It is possible to train the model from different initial random network configurations. The
ensemble of several neural networks has been shown to perform better than a single network.
However, note that the time required to train a model increases linearly with this
parameter.
Number of hidden neurons:
A higher number of hidden neurons in the ANNs potentially allows detecting higher order correlations, but increases the number of parameters of the model. Different hidden layer sizes can be specified in a comma separated list (e.g. 3,7,12,20), in which case an
ensemble of networks with different architectures is constructed.
Amino acid encoding:
Amino acids must be converted to numbers in order to be presented to the neural networks. Sparse encoding converts an amino acid into a binary vector, whereas Blosum encoding uses the BLOSUM62 substitution scores, accounting for physicochemical similarity between amino acids. Choosing the "Combined" option, networks are trained both with Blosum and Sparse encoding, combining the predictions of the two approaches. Note that Blosum encoding is only available if the training data uses the standard one-letter 20 amino acids alphabet (used as default).
Maximum length for Deletions:
Allow deletions in the alignment. For a description of how deletions (and insertions) are treated refer to this paper.
Maximum length for Insertions:
Allow insertions in the alignment.
Only allow insertions in sequences shorter than the motif length:
Apply insertions in any sequence, or only on sequences shorter than the motif length.
Burn-in period:
The burn-in is a number of initial iterations where no deletions or insertions are allowed. As gaps increase dramatically the number of possible solutions, it may be useful to use a burn-in period > 0 in order to limit the search space in the initial training phases.
Impose amino acid preference at P1 during burn-in:
In some cases, you may have some prior knowledge of the expected binding motif. For example, most HLA-DR molecules have a preference for hydrophobic residues (ILVMFYW) at P1. Such prior knowledge can help guiding the networks towards finding the correct binding core, and is applied only in the very first few iterations (specificed by the burn-in parameter). After the burn-in iterations, the restraint on the specified residues at P1 is removed, and the networks will consider all possible binding cores.
Preferred residues at P1:
Together with the previous option (impose amino acid preference at P1), this parameters allows specifying which subset of residues should be preferred at P1 of the binding core during the burn-in iterations.
Length of the PFR for composition encoding:
In some instances, the amino acid composition of the regions surrounding the motif core (peptide flanking region, PFR) can have an influence on the response. See for example in this paper, where the amino acid composition of a PFR of at least two amino acids around the core was shown to influence peptide-MHC binding strength. With this option you can specify the length of the regions flanking the alignment core, which will be encoded as input to NNAlign.
Encode PFR composition as sparse:
By default, the composition of the regions flanking the binding cores is encoded using the Blosum substitution matrix. Turning this option on, the raw frequency of each amino acid in the PFR (sparse alphabet) is used for encoding.
Encode PFR length:
Encodes the length of the flanks, i.e. the number of amino acids before/after the motif core. It essentially bears information about the position of the core within the peptide, if it is found at the extremes or in the middle. If this option is set to N > 0, the flank length is truncated to N amino acids, if N = 0 the encoding is unbounded (recommended). Setting this option to -1 disables the encoding.
Expected peptide length for encoding:
Assigns input neurons to encode the length of the input sequences if set to > 0. For an optimal encoding, give a rough estimate of the expected optimal peptide length.
Binned peptide length encoding:
Encode peptide length with individual input neurons for different lengths. For example, setting this parameter to "8,9,10,11" will create four separate input neurons, each activated only by peptides of the corresponding length (<=8, 9, 10, >=11 respectively). This encoding is different from the option "expected peptide length for encoding" which uses a continuous function that interpolates the possible peptide lengths.
Load receptor pseudo-sequences:
If you have different receptors associated with your training examples, specify the receptor names as the third column in the training file. Then, upload here a file with two columns: the receptor names in the first, and the aligned pseudo-sequences in the second. Note that pseudo-sequences must all have the same length and be in the same alphabet as the training sequences (including X).
Number of networks (per fold) in the final ensemble:
When training with cross-validation, each neural network's performance is evaluated
in terms of Pearson's correlation between target and predicted values. The
top N networks (for each cross-validation fold) can be selected using this
parameter, and they will constitute the final model.
Sort results by prediction value:
Predictions can be sorted by the NNAlign predicted value. If left unticked, sequences are presented in their original order.
Exclude offset correction:
Offset correction is a procedure that realigns individual networks to enhance the combined sequence motif (see the section "Improving the LOGO sequence motif representation by an offset correction" in this paper. You can disable offset correction by ticking this box.
Show all logos in the final ensemble:
Displays the sequence motif identified by each neural network in the model.
Length of peptides generated from FASTA entries:
Evaluation data submitted in FASTA format will be digested into fragments of the specified length. These peptides will then be submitted to the network ensemble to scan for the presence of the learned sequence motifs.
Sort evaluation results by prediction value:
Predictions on the independent evaluation set can be sorted by the NNAlign predicted value. If left unticked, sequences are presented in their original order.
Threshold on evaluation set predictions:
For large FASTA file submissions, the size of the results file may become very big. Use this parameter to limit the size of evaluation set results, and only show sequences with high predicted values. It should be given as a number between 0 and 1 (set to 0, all results will be displayed).
At any time during the wait you may enter your e-mail address and leave
the window. Your job will continue and you will be notified by e-mail when it has
terminated. The e-mail message will contain the URL under which the results are
stored; they will remain on the server for 24 hours for you to collect them.
|
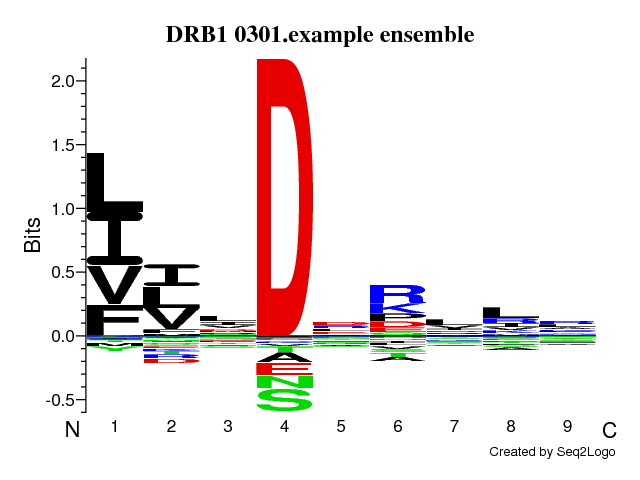
Version: 2.1 Run ID: 12857 Run Name: DRB1_0301.example Training dataRead 1715 unique sequencesView data distribution (See Instructions for optimal data distribution) Pre-processing: Linear rescale |
Neural network architectureMotif length: 9Flanking region (PFR) size: 3 Number of hidden neurons: 5,15 Peptide length encoding: 13 Flank length encoding: 0 Maximum length of deletions in alignment: 0 Maximum length of insertions in alignment: 0 Amino acid numerical encoding: Blosum Number of training cycles: 500 Number of NN seeds: 4 Number of networks in final ensemble: 40 Stop training on best test-set performance: Yes Cross-validation setup: Simple Folds for cross-validation : 5 Method to create subsets: Random |
NNAlign: a platform to construct and evaluate artificial neural network models of receptor-ligand interactions
Morten Nielsen1,2,
Massimo Andreatta1
Nucleic Acids Research, 2017 Apr 12. doi: 10.1093/nar/gkx276
1Instituto de Investigaciones Biotecnológicas, Universidad Nacional de San Martín, 1650 San Martín, Argentina
Peptides are extensively used to characterize functional or (linear) structural aspects of receptor-ligand interactions in biological systems e.g. SH2, SH3, PDZ peptide-recognition domains, the MHC membrane receptors and enzymes such as kinases and phosphatases. NNAlign is a method for the identification of such linear motifs in biological sequences. The algorithm aligns the amino acid or nucleotide sequences provided as training set, and generates a model of the sequence motif detected in the data. The webserver allows setting up cross-validation experiments to estimate the performance of the model, as well as evaluations on independent data. Many features of the training sequences can be encoded as input, and the network architecture is highly customizable. The results returned by the server include a graphical representation of the motif identified by the method, performance values and a downloadable model that can be applied to scan protein sequences for occurrence of the motif. While its performance for the characterization of peptide-MHC interactions is widely documented, we extended NNAlign to be applicable to other receptor-ligand systems as well. Version 2.0 supports alignments with insertions and deletions, encoding of receptor pseudo-sequences, and custom alphabets for the training sequences. The server is available at http://www.cbs.dtu.dk/services/NNAlign-2.0
2Department of Bio and Health Informatics, Technical University of Denmark, DK-2800 Lyngby, Denmark
Please click on the version number to activate the corresponding server.
| 2.1 |
Online since 01/11/2017 Implements "binned" peptide length encoding, and preferred P1 residues during burn-in iterations |
| 2.0 |
Online since 02/09/2016 Updates from the previous version include:
|
| 1.4 | Uses an improved algorithm for offset correction, that allows alignment of PSSMs on a window size of the same length as the core length (online since 01/12/2011). |
| 1.3 | Enhanced interface and visualization of LOGOs, and statistics for all motif lengths in an interval. Added Log-odds matrix representation of the motif (online since 23/06/2011). |
| 1.2 | Includes re-aligment of network cores for a better logo representation (online since 20/03/2011). |
| 1.1 | Implements several new options and fixes some bugs of version 1.0 (online since 25/01/2011). |
| 1.0 | (online since 06/01/2011). |
If you need help regarding technical issues (e.g. errors or missing results) contact Technical Support. Please include the name of the service and version (e.g. NetPhos-4.0) and the options you have selected. If the error occurs after the job has started running, please include the JOB ID (the long code that you see while the job is running).
If you have scientific questions (e.g. how the method works or how to interpret results), contact Correspondence.
Correspondence:
Technical Support: