DTU Health Tech
Department of Health Technology
This link is for the general contact of the DTU Health Tech institute.
If you need help with the bioinformatics programs, see the "Getting Help" section below the program.
DTU Health Tech
Department of Health Technology
This link is for the general contact of the DTU Health Tech institute.
If you need help with the bioinformatics programs, see the "Getting Help" section below the program.
DeepLoc 2.0 predicts the subcellular localization(s) of eukaryotic proteins. DeepLoc 2.0 is a multi-label predictor, which means that is able to predict one or more localizations for any given protein. It can differentiate between 10 different localizations: Nucleus, Cytoplasm, Extracellular, Mitochondrion, Cell membrane, Endoplasmic reticulum, Chloroplast, Golgi apparatus, Lysosome/Vacuole and Peroxisome. Additionally, DeepLoc 2.0 can predict the presence of the sorting signal(s) that had an influence on the prediction of the subcellular localization(s).
Prokaryotic proteins: To predict the locations of proteins in prokaryotes, use
DeepLocPro.
RNA: To predict the locations of RNA, use DeepLocRNA.
| NOTE: This is not the newest version of DeepLoc. To use the current version with the added capability of predicting membrane protein types, please go to DeepLoc 2.1! |
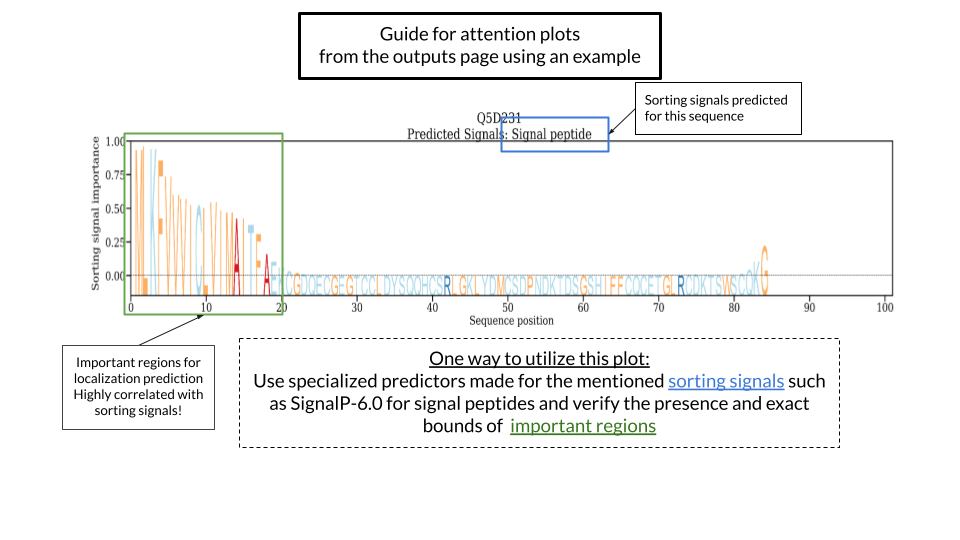
The DeepLoc 2.0 server predicts the multi-label subcellular localization of eukaryotics proteins using Neural Networks algorithm trained on Uniprot proteins with experimental evidence of subcellular localization. The model can predict whether a protein can be in one or multiple localizations inside the eukaryotic cell. It only uses the sequence information to perform the prediction. Additionally, DeepLoc 2.0 can predict the presence of the sorting signal(s) that had an influence on the prediction of the subcellular localization(s). The importance of each amino acid in the predicted localization is also included as an "attention" plot. Positions in the sequence with a high attention value are deemed more relevant for the prediction. This does not mean that a particular amino acid is very important for the prediction but that a region in the neighbourhood of those positions has more weight in the final prediction of the model.
The DeepLoc 2.0 server can be run using two versions of the same model.
The DeepLoc 2.0 server requires protein sequence(s) in fasta format, and can not handle nucleic acid sequences.
Two different versions of the output can be selected before running DeepLoc 2.0. The long output will generate an attention plot per sequence while the short output will not generate any plots.
Paste protein sequence(s) in fasta format or upload a fasta file.
After the server successfully finishes the job, a summary page shows up. If an error happens during the prediction a log will appear specifying the error.
The DeepLoc 2.0 output is composed of three main components:
The dataset used to train and test the DeepLoc 2.0 server is available here:
The Partition column in the Training/Validation set indicates the five partitions (0-4) that the dataset was homology partitioned (maximum 30% sequence similarity).| 2.0 |
The current server. New in this version:
DeepLoc 2.0: multi-label subcellular localization prediction using protein language models. Vineet Thumuluri, Jose Juan Almagro Armenteros, Alexander Rosenberg Johansen, Henrik Nielsen, Ole Winther. Nucleic Acids Research, Web server issue 2022. |
| 1.0 |
The original DeepLoc server. Publication: DeepLoc: prediction of protein subcellular localization using deep learning Jose Juan Almagro Armenteros, Casper Kaae Sønderby, Søren Kaae Sønderby, Henrik Nielsen, Ole Winther. Bioinformatics, 33:3387–3395 (2017). |
If you need help regarding technical issues (e.g. errors or missing results) contact Technical Support. Please include the name of the service and version (e.g. NetPhos-4.0) and the options you have selected. If the error occurs after the job has started running, please include the JOB ID (the long code that you see while the job is running).
If you have scientific questions (e.g. how the method works or how to interpret results), contact Correspondence.
Correspondence:
Technical Support: